Description
This tool comprises of a compartmental stochastic model at a population-scale, which is based on three different components: entomological dynamics, transmission dynamics in the human population, and (optionally) spatial dynamics. Mosquitoes and humans reside in patches where their respective densities are assumed to be homogeneous. Humans may move between patches, propagating viruses spatially if infected or infectious. The mosquito dynamics is rainfall and temperature driven, and the transmission is temperature driven. The user may experiment with vector control affecting immature or mature mosquito stages, and vary virus importation in humans.Manual
Download the manualModel code (R)
Download the model code For experienced R users who want to run the model on their own computers, the R code of this shiny app is made available for download.Workflow
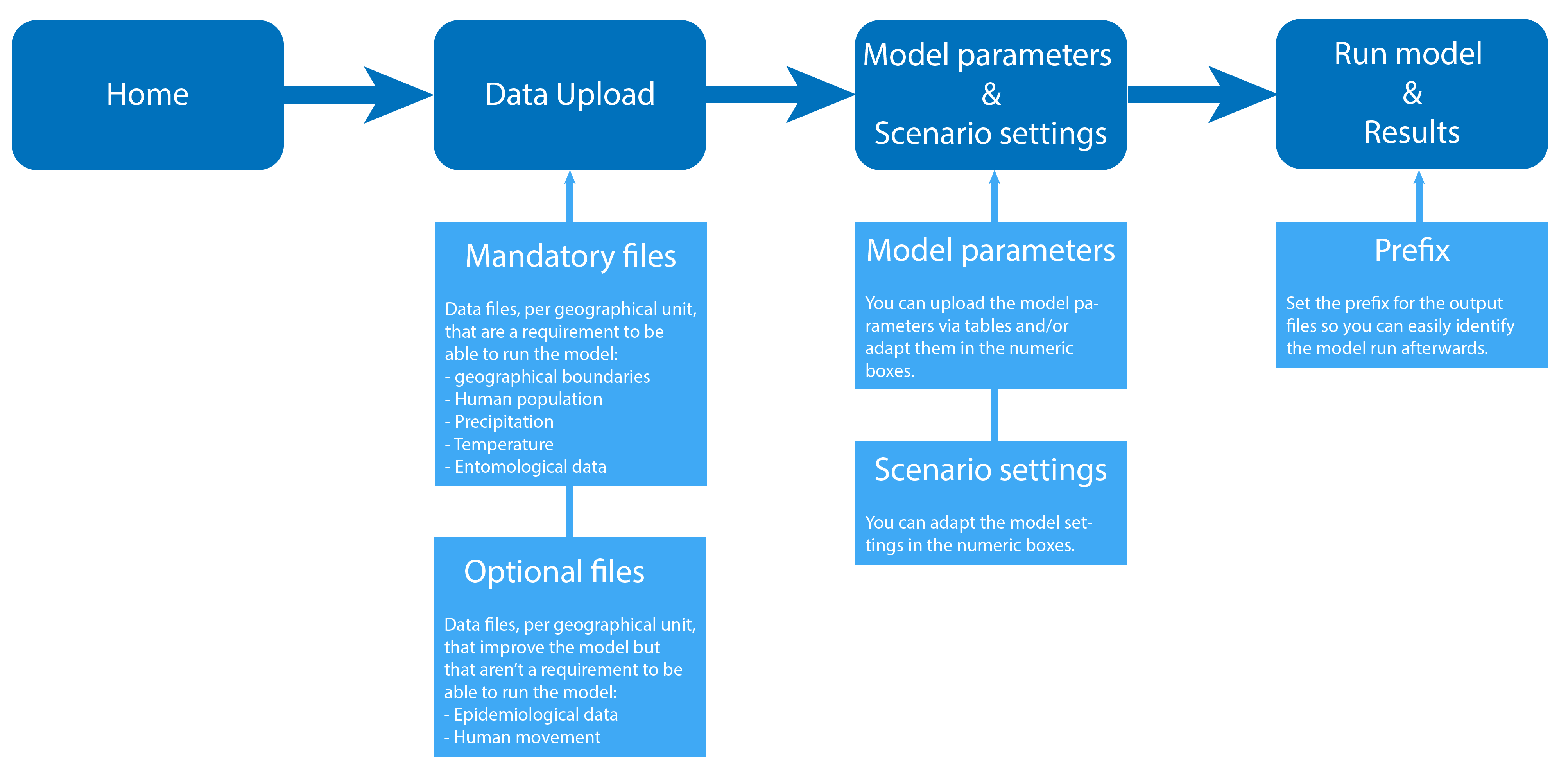
For a more detailed description please refer to the tool's manual.